| Record Information |
|---|
| Version | 1.0 |
|---|
| Creation Date | 2020-03-10 17:00:16 UTC |
|---|
| Update Date | 2026-03-26 16:20:27 UTC |
|---|
| BMDB ID | BMDB0096043 |
|---|
| Secondary Accession Numbers | |
|---|
| Metabolite Identification |
|---|
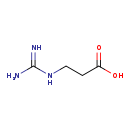
| Common Name | Beta-Guanidinopropionic acid |
|---|
| Description | Guanidinopropionic acid (GPA), also known as guanidinopropionate or 3-guanidinopropionic acid or beta-guanidinopropionic acid, belongs to the class of organic compounds known as guanidines. Guanidines are compounds containing a guanidine moiety, with the general structure (R1R2N)(R3R4N)C=N-R5. GPA (C₄H₉N₃O₂) is structurally related to creatine that has been detected at low levels in mammalian tissues including blood, brain, liver, kidney, and urine, suggesting a minor endogenous origin possibly arising through transamidination reactions between arginine and β-alanine occurring in the kidney (PMID: 1312834 ; PMID: 33890206 ). Functionally, GPA acts as a competitive inhibitor of creatine transport, entering cells via the creatine transporter and reducing intracellular creatine and phosphocreatine levels, thereby disrupting the creatine kinase energy-buffering system. This induces a state of cellular energy stress, activating adaptive pathways such as AMP-activated protein kinase and promoting shifts toward oxidative metabolism, which has led to its widespread use as an experimental tool to study muscle energetics, mitochondrial function, and metabolic regulation (PMID: 33890206 ). In animal models, GPA administration has been associated with decreased phosphocreatine levels in muscle and heart, improved insulin sensitivity, reduced adiposity, and alterations in brown adipose tissue metabolism, although chronic exposure can impair growth and muscle function. GPA is typically synthesized chemically through reactions that introduce a guanidino group onto a propionic acid backbone using guanidine derivatives and activated intermediates. While GPA does not play a major essential physiological role in humans, it is increasingly recognized as a guanidino compound that can accumulate under conditions of impaired renal function, where it may contribute to the pool of uremic toxins (PMID: 3960241 ). In this context, elevated GPA levels may exacerbate metabolic disturbances by interfering with cellular energy homeostasis and contributing to mitochondrial dysfunction, highlighting its potential relevance in chronic kidney disease and related metabolic complications. |
|---|
| Structure | |
|---|
| Synonyms | | Value | Source |
|---|
| 3-Guanidinopropanoate | ChEBI | | beta-GPA | ChEBI | | N-[Amino(imino)methyl]-beta-alanine | ChEBI | | Guanidinopropionic acid | Kegg | | 3-Guanidinopropanoic acid | Generator | | b-GPA | Generator | | Β-gpa | Generator | | N-[Amino(imino)methyl]-b-alanine | Generator | | N-[Amino(imino)methyl]-β-alanine | Generator | | Guanidinopropionate | Generator | | b-Guanidinopropionate | Generator | | b-Guanidinopropionic acid | Generator | | beta-Guanidinopropionate | Generator | | Β-guanidinopropionate | Generator | | Β-guanidinopropionic acid | Generator | | 3-Guanidinopropionic acid | HMDB | | beta-Guanadinopropionate | HMDB | | beta-Guanidinopropionic acid. | HMDB | | beta-Guanidine propionic acid | MeSH, HMDB | | Guanidine propionate | MeSH, HMDB | | beta-Guanidinopropionic acid | ChEBI | | Amidino beta-alanine | MeSH |
|
|---|
| Chemical Formula | C4H9N3O2 |
|---|
| Average Molecular Weight | 131.1332 |
|---|
| Monoisotopic Molecular Weight | 131.069476547 |
|---|
| IUPAC Name | 3-carbamimidamidopropanoic acid |
|---|
| Traditional Name | 3-guanidinopropanoic acid |
|---|
| CAS Registry Number | Not Available |
|---|
| SMILES | NC(=N)NCCC(O)=O |
|---|
| InChI Identifier | InChI=1S/C4H9N3O2/c5-4(6)7-2-1-3(8)9/h1-2H2,(H,8,9)(H4,5,6,7) |
|---|
| InChI Key | KMXXSJLYVJEBHI-UHFFFAOYSA-N |
|---|
| Chemical Taxonomy |
|---|
| Description | belongs to the class of organic compounds known as guanidines. Guanidines are compounds containing a guanidine moiety, with the general structure (R1R2N)(R3R4N)C=N-R5. |
|---|
| Kingdom | Organic compounds |
|---|
| Super Class | Organic nitrogen compounds |
|---|
| Class | Organonitrogen compounds |
|---|
| Sub Class | Guanidines |
|---|
| Direct Parent | Guanidines |
|---|
| Alternative Parents | |
|---|
| Substituents | - Guanidine
- Organic 1,3-dipolar compound
- Propargyl-type 1,3-dipolar organic compound
- Carboximidamide
- Monocarboxylic acid or derivatives
- Carboxylic acid
- Carboxylic acid derivative
- Organic oxygen compound
- Organopnictogen compound
- Organic oxide
- Hydrocarbon derivative
- Organooxygen compound
- Carbonyl group
- Aliphatic acyclic compound
|
|---|
| Molecular Framework | Aliphatic acyclic compounds |
|---|
| External Descriptors | |
|---|
| Ontology |
|---|
| Status | Expected but not Quantified |
|---|
| Origin | |
|---|
| Biofunction | Not Available |
|---|
| Application | Not Available |
|---|
| Cellular locations | Not Available |
|---|
| Physical Properties |
|---|
| State | Not Available |
|---|
| Experimental Properties | | Property | Value | Reference |
|---|
| Melting Point | Not Available | Not Available | | Boiling Point | Not Available | Not Available | | Water Solubility | Not Available | Not Available | | LogP | Not Available | Not Available |
|
|---|
| Predicted Properties | |
|---|
| Spectra |
|---|
| Spectra | | Spectrum Type | Description | Splash Key | |
|---|
| Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (Non-derivatized) - 70eV, Positive | splash10-0076-9100000000-351fe5459bc2aa982f1d | View in MoNA |
|---|
| Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (1 TMS) - 70eV, Positive | splash10-00dl-9700000000-8fa57b59d32757572966 | View in MoNA |
|---|
| Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (Non-derivatized) - 70eV, Positive | Not Available | View in JSpectraViewer |
|---|
| LC-MS/MS | LC-MS/MS Spectrum - LC-ESI-QTOF , negative | splash10-000i-9000000000-4458e96907418a84d54b | View in MoNA |
|---|
| LC-MS/MS | LC-MS/MS Spectrum - LC-ESI-QQ , positive | splash10-001i-0900000000-555c4102f4a65d8aed9b | View in MoNA |
|---|
| LC-MS/MS | LC-MS/MS Spectrum - LC-ESI-QQ , positive | splash10-00di-9400000000-202f65d00956e051d5b0 | View in MoNA |
|---|
| LC-MS/MS | LC-MS/MS Spectrum - LC-ESI-QQ , positive | splash10-00di-9000000000-2ba68363493ccaf36ede | View in MoNA |
|---|
| LC-MS/MS | LC-MS/MS Spectrum - LC-ESI-QQ , positive | splash10-05fr-9000000000-245d3980a77046e1db8d | View in MoNA |
|---|
| LC-MS/MS | LC-MS/MS Spectrum - LC-ESI-QQ , positive | splash10-052f-9000000000-4659c75c5d7116058d41 | View in MoNA |
|---|
| LC-MS/MS | LC-MS/MS Spectrum - LC-ESI-QTOF , positive | splash10-0089-6900000000-23e0505168600ceb154d | View in MoNA |
|---|
| LC-MS/MS | LC-MS/MS Spectrum - 35V, Positive | splash10-03k9-9000000000-04ea2687312454f3f43b | View in MoNA |
|---|
| Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Positive | splash10-03di-2900000000-d05e3c3a923d5ffaf398 | View in MoNA |
|---|
| Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Positive | splash10-03k9-9200000000-6b780b0080ec3851a2a1 | View in MoNA |
|---|
| Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Positive | splash10-03fu-9000000000-585f46f520a38f066ad0 | View in MoNA |
|---|
| Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Negative | splash10-001r-9600000000-e7ba0696feae67ecd2c1 | View in MoNA |
|---|
| Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Negative | splash10-059l-9000000000-1ad5c9a7f49f2b5eab63 | View in MoNA |
|---|
| Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Negative | splash10-0006-9000000000-28676e3aacf9b4d8ea55 | View in MoNA |
|---|
| Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Negative | splash10-000i-9000000000-d58857401b43d8ccc8b5 | View in MoNA |
|---|
| Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Negative | splash10-006x-9000000000-5900927063e733df75f2 | View in MoNA |
|---|
| Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Negative | splash10-0006-9000000000-1c8520c5f7efb1187e43 | View in MoNA |
|---|
| Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Positive | splash10-02tc-7900000000-23251cef83c81d03c926 | View in MoNA |
|---|
| Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Positive | splash10-00di-9000000000-9b84fcc1808b7ece8370 | View in MoNA |
|---|
| Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Positive | splash10-0006-9000000000-dd976c4f7e1faaec59fd | View in MoNA |
|---|
| 1D NMR | 13C NMR Spectrum (1D, 100 MHz, D2O, predicted) | Not Available | View in JSpectraViewer |
|---|
| 1D NMR | 1H NMR Spectrum (1D, 100 MHz, D2O, predicted) | Not Available | View in JSpectraViewer |
|---|
| 1D NMR | 13C NMR Spectrum (1D, 1000 MHz, D2O, predicted) | Not Available | View in JSpectraViewer |
|---|
| 1D NMR | 1H NMR Spectrum (1D, 1000 MHz, D2O, predicted) | Not Available | View in JSpectraViewer |
|---|
| 1D NMR | 13C NMR Spectrum (1D, 200 MHz, D2O, predicted) | Not Available | View in JSpectraViewer |
|---|
| 1D NMR | 1H NMR Spectrum (1D, 200 MHz, D2O, predicted) | Not Available | View in JSpectraViewer |
|---|
| 1D NMR | 13C NMR Spectrum (1D, 300 MHz, D2O, predicted) | Not Available | View in JSpectraViewer |
|---|
| 1D NMR | 1H NMR Spectrum (1D, 300 MHz, D2O, predicted) | Not Available | View in JSpectraViewer |
|---|
| 1D NMR | 13C NMR Spectrum (1D, 400 MHz, D2O, predicted) | Not Available | View in JSpectraViewer |
|---|
| 1D NMR | 1H NMR Spectrum (1D, 400 MHz, D2O, predicted) | Not Available | View in JSpectraViewer |
|---|
| 1D NMR | 13C NMR Spectrum (1D, 500 MHz, D2O, predicted) | Not Available | View in JSpectraViewer |
|---|
| 1D NMR | 1H NMR Spectrum (1D, 500 MHz, D2O, predicted) | Not Available | View in JSpectraViewer |
|---|
| 1D NMR | 13C NMR Spectrum (1D, 600 MHz, D2O, predicted) | Not Available | View in JSpectraViewer |
|---|
| 1D NMR | 1H NMR Spectrum (1D, 600 MHz, D2O, predicted) | Not Available | View in JSpectraViewer |
|---|
| 1D NMR | 13C NMR Spectrum (1D, 700 MHz, D2O, predicted) | Not Available | View in JSpectraViewer |
|---|
| 1D NMR | 1H NMR Spectrum (1D, 700 MHz, D2O, predicted) | Not Available | View in JSpectraViewer |
|---|
| 1D NMR | 13C NMR Spectrum (1D, 800 MHz, D2O, predicted) | Not Available | View in JSpectraViewer |
|---|
| 1D NMR | 1H NMR Spectrum (1D, 800 MHz, D2O, predicted) | Not Available | View in JSpectraViewer |
|---|
| 1D NMR | 13C NMR Spectrum (1D, 900 MHz, D2O, predicted) | Not Available | View in JSpectraViewer |
|---|
| 1D NMR | 1H NMR Spectrum (1D, 900 MHz, D2O, predicted) | Not Available | View in JSpectraViewer |
|---|
|
|---|
| General References | - Thomas G, Ramwell PW: Identification of guanidino succinate as a putative endogenous source of the endothelium derived relaxing factor. Biochem Biophys Res Commun. 1992 Mar 16;183(2):584-9. doi: 10.1016/0006-291x(92)90522-m. [PubMed:1312834 ]
- Dorigatti JD, Thyne KM, Ginsburg BC, Salmon AB: Beta-guanidinopropionic acid has age-specific effects on markers of health and function in mice. Geroscience. 2021 Jun;43(3):1497-1511. doi: 10.1007/s11357-021-00372-8. Epub 2021 Apr 23. [PubMed:33890206 ]
- Gurreri G, Ghiggeri G, Salvidio G, Garibotto G, Robaudo C, Deferrari G: Effects of hemodialysis on guanidinopropionic acid metabolism. Nephron. 1986;42(4):295-7. doi: 10.1159/000183691. [PubMed:3960241 ]
|
|---|